Early History of Epigenetics
Although a relatively new field, epigenetics has a long and storied history dating back to the mid 18th century. Like many principles in science, epigenetics started with the recognition of an inexplicable phenomenon – that is, how are traits propagated between generations? The first seeds of epigenetics were arguably planted by Charles Darwin, the 19th century biologist and father of evolutionary theory. Darwin argued for a mechanism of heredity termed pangenesis.
In this model, indeterminate molecules referred to as “gemmules” were released from cells and transported via the bloodstream to one’s germ cells. These proposed hereditary units were thought to possess plasticity in the sense that different kinds of gemmules would be produced depending on environmental conditions. Darwin’s idea formalized a notion, often misattributed to Jean-Baptiste Lamarck, that organism’s can pass on traits that develop as a product of their lifestyle. (The origin of this concept – “inheritance of acquired characteristics” – is beyond the scope of this review). A commonly cited example is that of the giraffe. Since giraffes extend their necks up to high trees in search for food, Lamarck proposed that successive generations which reached for higher and higher trees would pass on increasingly elaborate necks.

A Lamarckian idea that the act of stretching one’s neck could lead to change in phenotype across generations.
Early 20th Century Epigenetic History
Until the mid-twentieth century, it remained unclear how the fertilized egg develops into a fully functional human with processes compartmentalized into distinct organs. However, it was known that, in general, all cells in the body contained the same genome. That is to say, the sequence of A’s, T’s, C’s, and G’s were no different in one’s neurons than in one’s muscles. The question was: how can cells function so differently with the same genetic code? A series of informative experiments shed light on this mystery.
Although cells throughout the human body have identical DNA, they may have very different form and function between types. This is due to differences in epigenetics.
By transplanting nuclei from xenopus (frog) somatic cells into empty egg cells, Nobel Laureate John Gurdon paved the way for our understanding of how cell types are determined. This work definitively demonstrated the plasticity of cell identity. The embryos developed into adult frogs that were fertile and able to reproduce. Thus, the full set of processes necessary for embryogenesis were recapitulated in the egg cell that received the transplanted nucleus. This implied that the factors influencing cell fate decisions are localized to the nucleus. Given the supposed genetic homogeneity of these cells, it was clear that some other factor within the nucleus was at play.
John Gurdon’s experiment showing that taking the nucleus from an adult frog skin cell, or the gut cell from a tadpole, and inserting into a egg cell without a nucleus, the egg developed into a fertile tadpole.
The term epigenetics is literally translated as being above or upon genetics, which is befitting given that it is an extra layer of biological information overlaid on the genome. In practice, epigenetics is defined as changes to DNA excluding those which alter the actual nucleotide sequence. However, a great irony arose in 1953 when the double-helical structure of DNA was elucidated by James Watson and Francis Crick; The composition of DNA was characterized well after its chemical modifications (epigenetics) were. Indeed, and chromatin-mediated silencing of gene expression as outlined by Edgar Stedman preceded the DNA structure by 3 years. Histones were documented as far back as 1884 by Albrecht Kossel.
Conrad Waddington coined the term, “epigenetic landscape” and is thought of by many to be the father of epigenetics. Waddington’s landscape concept was a pivotal concept in developmental biology as it attempted to explain how a static set of DNA sequences could dynamically give rise to a complex organism. Pioneering work by Waddington also demonstrated compelling evidence for inheritance of a seemingly acquired characteristic in drosophila fruit flies.
Modern Epigenetics
Modern day epigenetics is comprised of methylation, histones and their modifications, and non-coding RNAs. Acting in concert, these features coordinate intricate regulatory mechanisms that tightly control the expression of the genome. The propensity of the environment for perturbing these epigenetic pathways has become increasingly evident. Contemporary scientists are therefore taking a fresh look at Lamarckian inheritance with the idea that epigenetics may be a validating manifestation of this phenomenon.
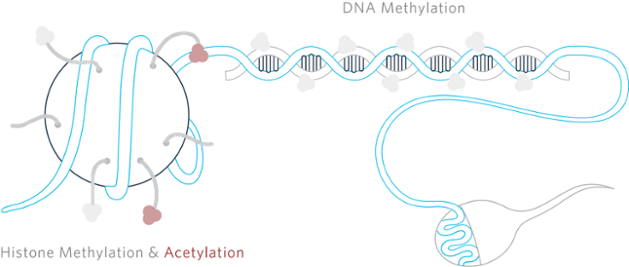
DNA Methylation
A tangible example of methylation is the inactivation of the second X chromosome in females which is systematically “shut down” during development. It wasn’t until 1975 that a mechanistic understanding of methylation began to take shape. In that year, using the X chromosome as an extreme example of epigenetic repression, Arthur Riggs proposed a silencing role for methylation which functioned to prevent the binding of proteins involved in gene regulation. This paradigm has since been refined, and already ten years later Adrian Bird made it known that methyl groups tend to preferentially inhabit CpG islands.
Histones and Histone Modification
Chromatin are the complex coiled structures formed through the interaction of DNA with associated histone proteins. These histones can undergo modification via acetylation, citrullination, methylation and phosphorylation of their amino residues. It is commonly accepted that loosening or tightening of the chromatin into euchromatin and heterochromatin respectively serves to modulate gene expression. This became clear with a landmark paper in 1964 by Alfrey, Faulkner and Mirsky that tied chromatin modifications to transcriptional regulation.
Non-Coding RNA
The term non-coding RNA and all of its subclasses refers to RNA that does not encode proteins. Unlike methylation, non-coding RNA is not a direct modification of DNA however, it merits falling under the category of epigenetics because it can regulate gene expression in the absence of a changing DNA sequence. RNA interference, a prominent subgroup of RNA interaction, was discovered by Andrew Fire and Craig Mello in 1998. They reasoned that these interfering RNAs could bind to messenger RNAs and neutralize them, thereby preventing translation into protein. To this day new types of non-coding RNA continue to be discovered with large intergenic non coding RNA (lincRNA) having only been discovered at the turn of the century.
The ability of the environment to establish epigenetic annotations has been extensively documented in recent years. In parallel, Lamarckian ideas have undergone a revival with a greater focus on rigorous mechanistic interrogation. As an illustrative example from 2014, prediabetes can change the sperm epigenome and predispose children to full-blown diabetes. This represents a clear link between lifestyle-induced non-genetic changes that end up reaching the offspring and dictating their health in a measurable way.
Cancer and Epigenetic Disease
Epigenetics has been implicated in the pathogenesis of diseases as diverse as cancer, neurodevelopmental disorders, autism, and fragile X. Once the role of epigenetics in normal biological processes became established, groups around the world began exploring its contribution to the etiology of disease. Cancer epigenetics, for example, has exploded with the recognition that methylation of promoter regions has a silencing effect on tumor suppressor genes which normally function to protect cells from turning cancerous. In 1983, Andrew Feinberg and Bert Vogelstein set the field ablaze when they began comparing the methylation profiles of known cancer genes between normal and malignant human tissue. In the case of cancer, these epigenetic changes are thought to arise within the lifespan of an organism. However, in other diseases, like Angelman syndrome, the abnormal epigenetic pattern is established before the child is born. Such a disorder is referred to as an imprinting disorder because the methylation occurs in the parents’ gametes and are passed on to their offspring. Angelman syndrome and the related Prader-Willi syndrome were the first disorders discovered to have an imprinting defect. It wasn’t until decades after the syndromes were first observed clinically that the associated chromosome 15 abnormalities were uncovered. Today, transgenerational inheritance of epigenetic disease states is an active area of research which has yielded surprising results. Thus far one thing is clear: there are an abundance of abnormal epigenetic changes which have the capacity to drive diverse pathological processes.
Male Factor Infertility and Epigenetics
Epigenetics has had a storied past in the scientific community, but its utility in the clinical realm is becoming ever more apparent. One disease application that looks particularly promising is that of male infertility diagnosis. Current male fertility tests are limited in the scope of parameters they can test leaving many cases of sub-fertility unexplained. There also exists an imbalance between the number of tests available for females versus males despite the fact that male factors comprise up to 50% of the couples infertility.Interestingly, a recent paper has suggested that there are methylation signatures of infertility in the sperm epigenome. Therefore, male infertility testing represents an important unmet clinical need.
In some cases of infertility, sperm DNA methylation may be altered. Thus, it is possible that DNA methylation may serve as a unique biomarker for male infertility.